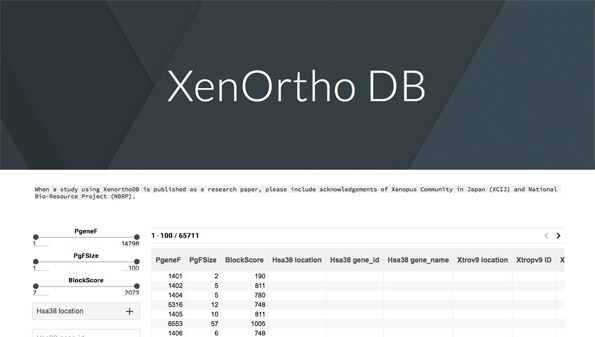
XenOrtho DB (external link) produced by Dr. Fukui (XCIJ, Chuo Univ.) and Dr. Igawa (NBRP X. tropicalis, Hiroshima Univ.)

When a study using XenOrthoDB is published as a research paper, please acknowledge the Xenopus Community in Japan (XCIJ) and National Bio-Resource Project (NBRP).
-
Japanese / English
XenOrtho DB (Xenopus Ortholog DataBase) is a database in which ortholog genes of Xenopus tropicalis, Xenopus laevis, and humans are listed.
Orthologs were identified through homology searches of genes in each pair of homologous synteny blocks (conserved regions within two sets of chromosomes). Synteny blocks were found with SyMAP4.0.
The release of this database is a project of NBRP X. tropicalis and greatly supported by Professor Fukui, Chuo University.
SyMAP 4.2 Tour (external link) is helpful in learning the terminology related to SyMap. If you have any other questions about XenOrtho DB, please contact Dr. Igawa (click “Contact Information” in the left side bar).
Toward application to medical and pharmacological research
The Xenopus-Human Ortholog Table
-
Xenopus vs. Human Orthologs
Copyright Ⓒ NBRP X. tropicalis. All rights reserved.